-
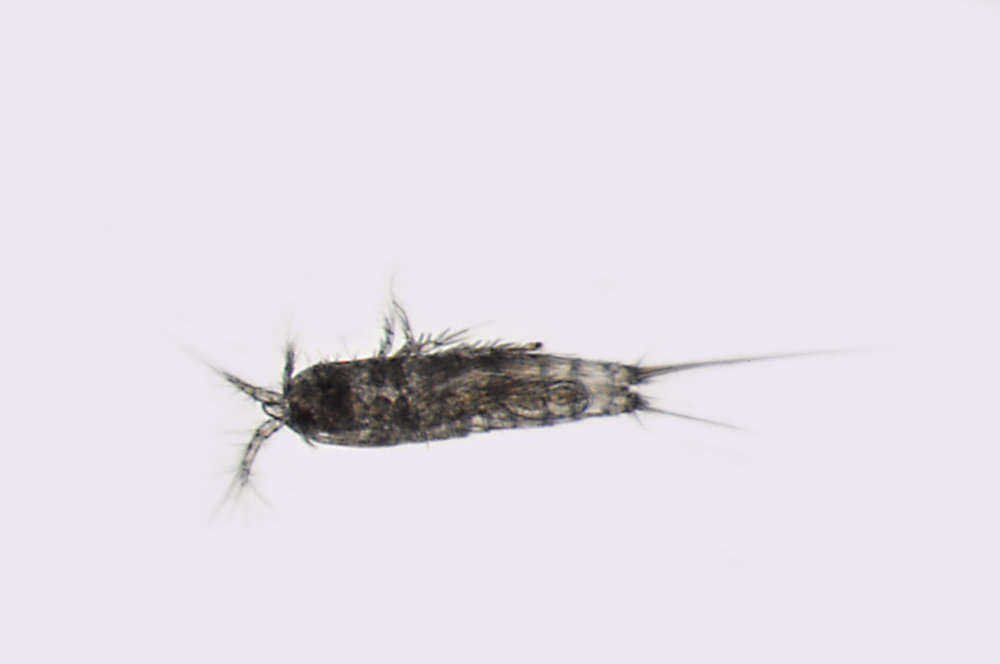
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
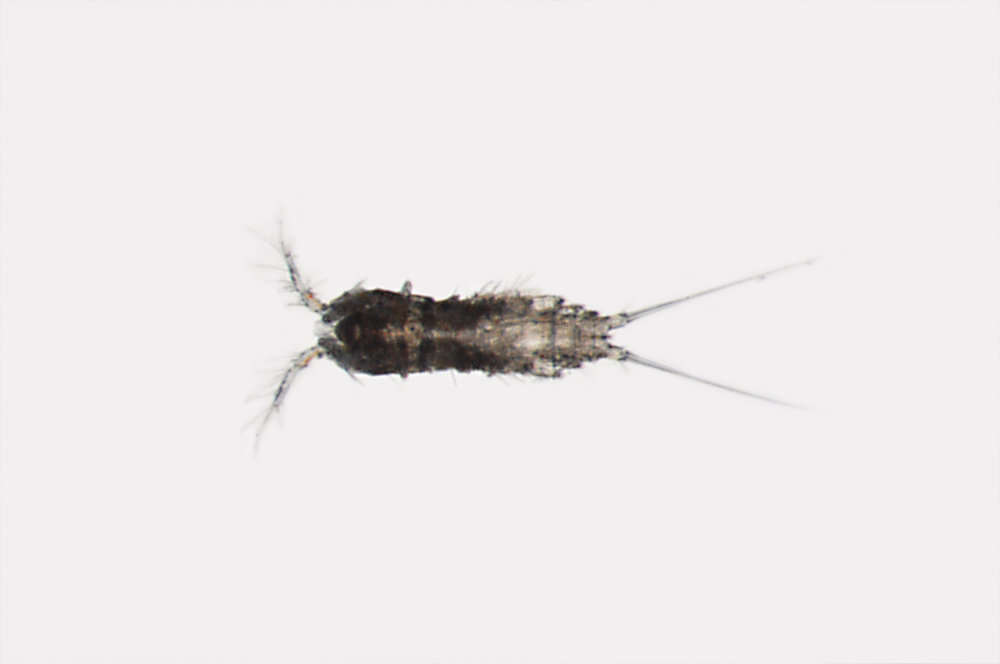
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
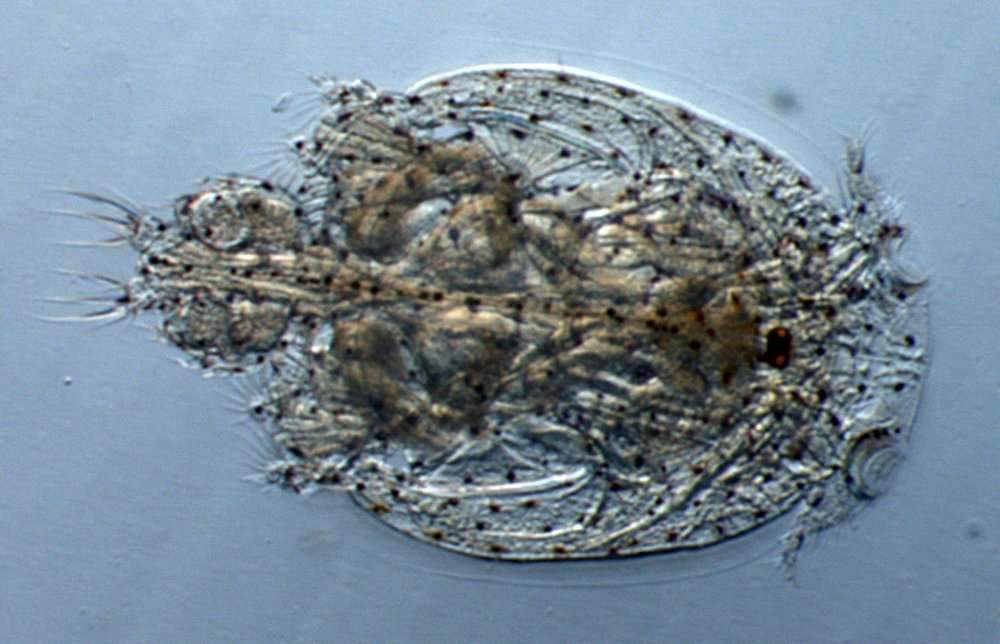
Tomislav Karanovic, Mark J. Grygier, Wonchoel Lee
Zookeys
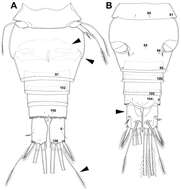
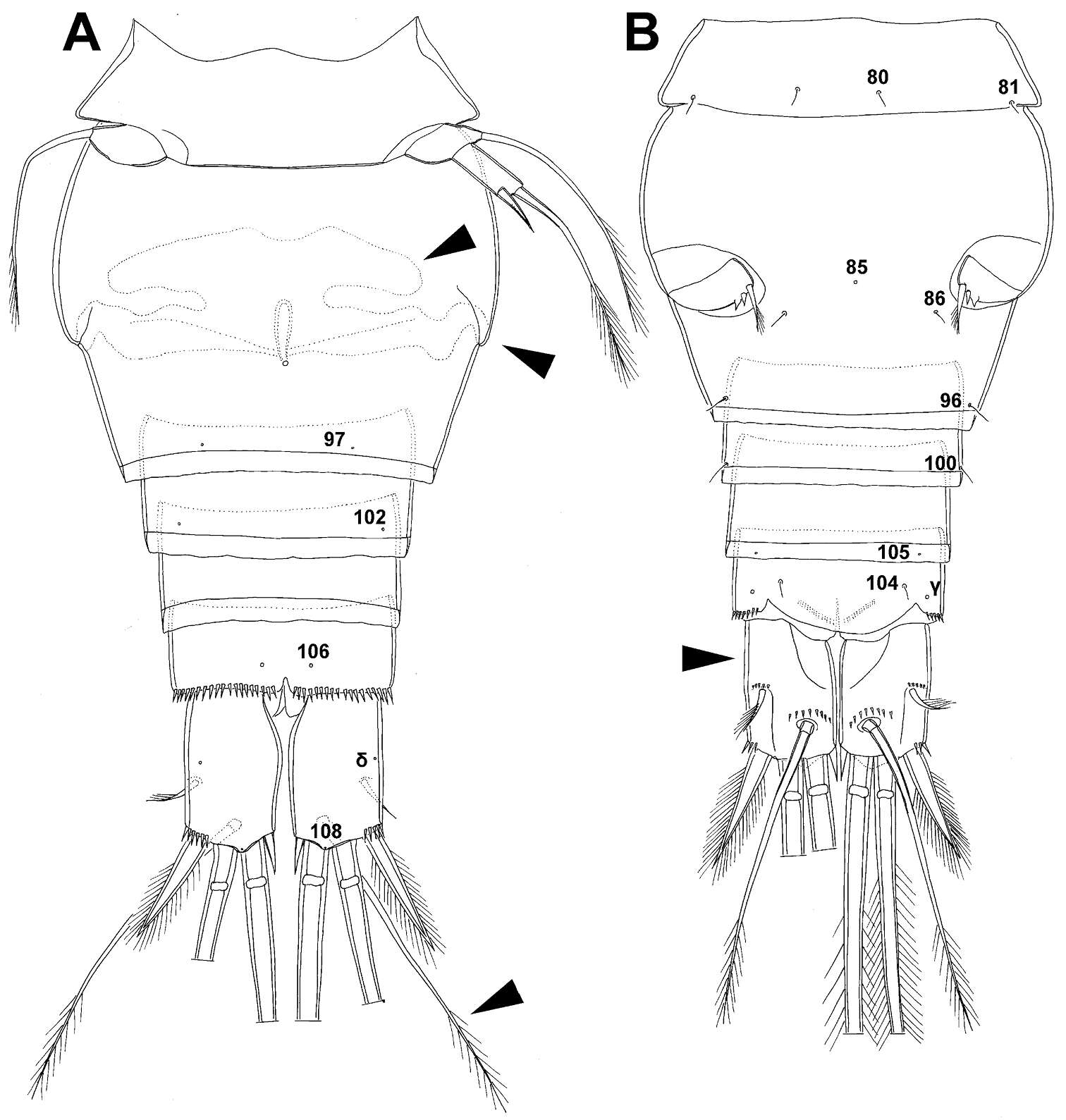
Figure 24.Diacyclops parahanguk sp. n., holotype female: A urosome, ventral view B urosome, dorsal view. Arabic numerals indicating sensilla and pores presumably homologous to those in Diacyclops ishidai sp. n. Greek letters indicating pores and sensilla homologous to those in Diacyclops hanguk sp. n. Arrows pointing most prominent specific features. Scale bar 100 μm.
-
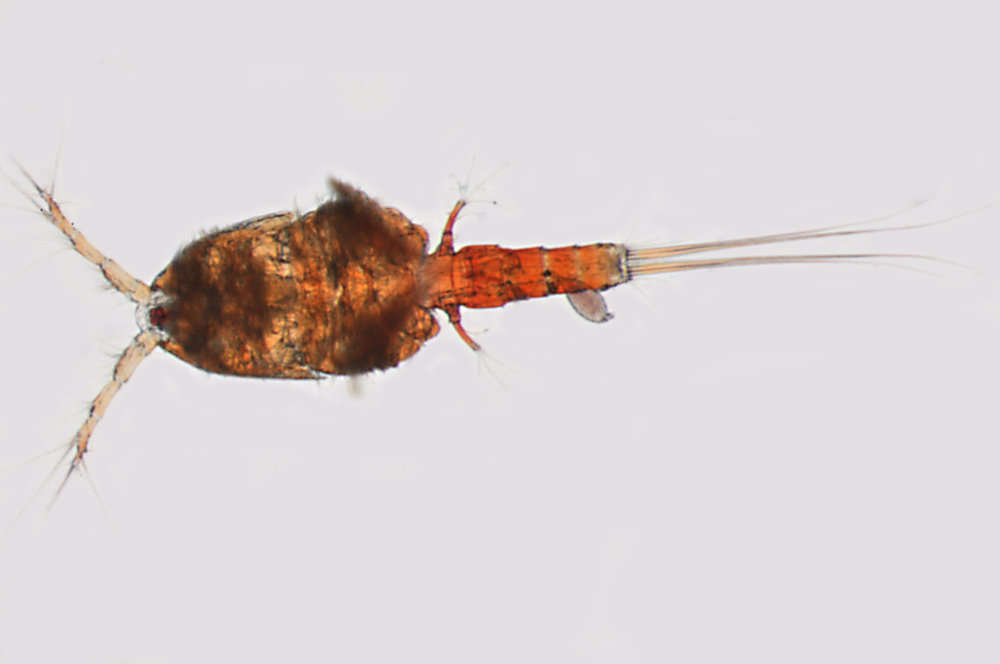
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
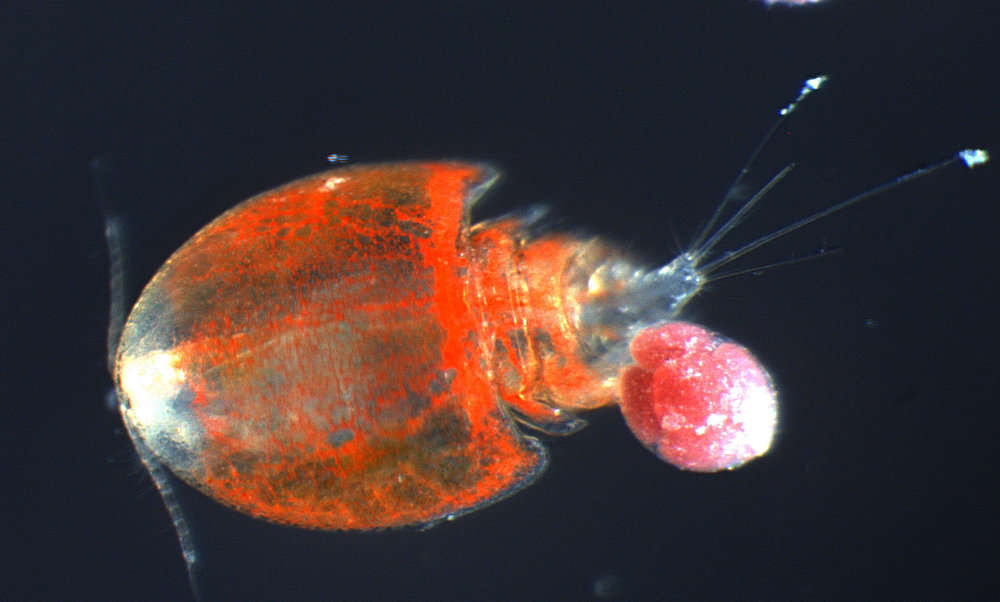
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
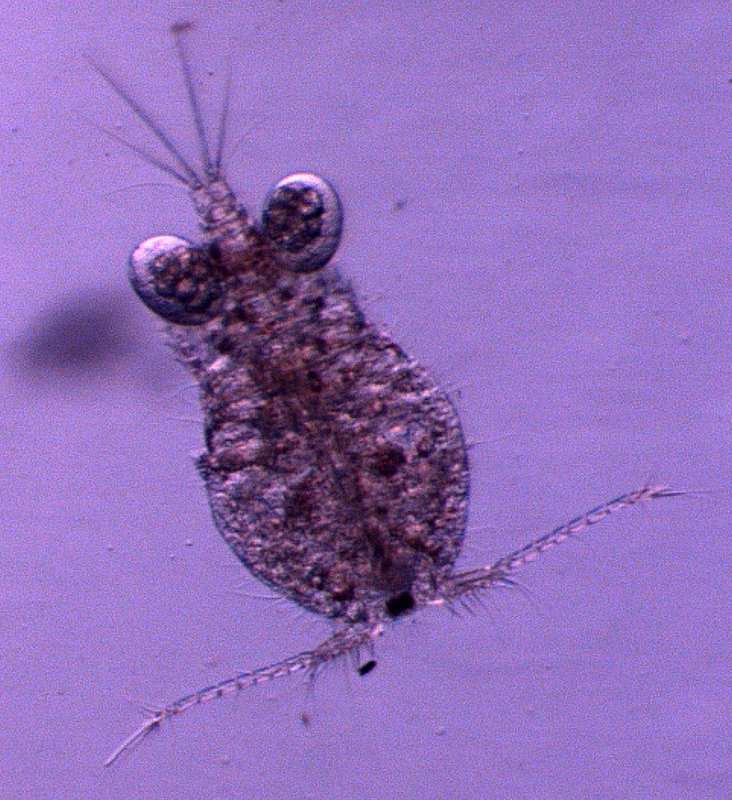
Tomislav Karanovic, Mark J. Grygier, Wonchoel Lee
Zookeys
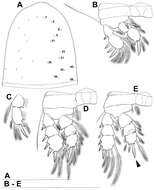
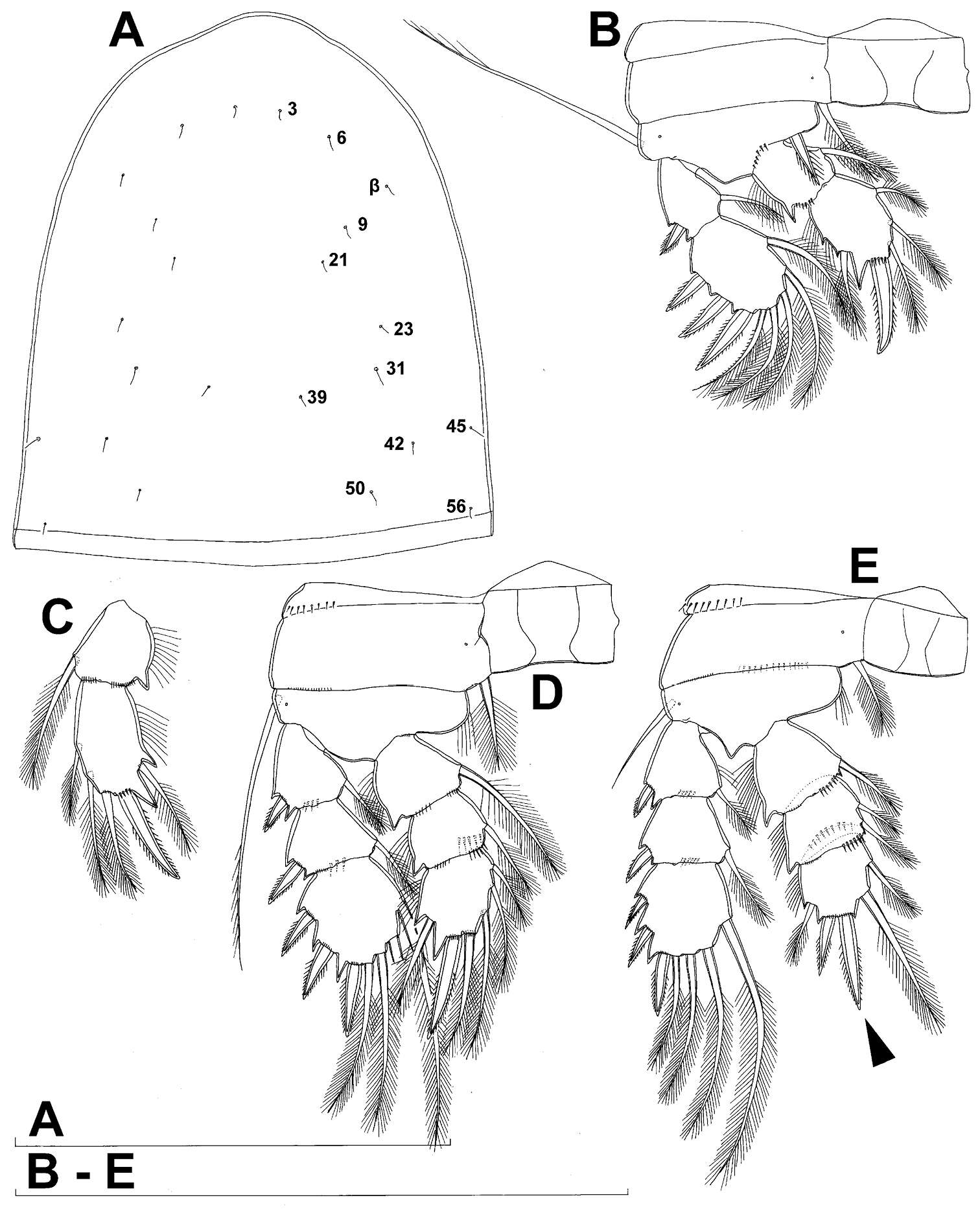
Figure 25.Diacyclops parahanguk sp. n., holotype female: A cephalothorax, dorsal view B first swimming leg, anterior view C endopod of second swimming leg, anterior view D third swimming leg, anterior view E fourth swimming leg, anterior view. Arabic numerals indicating sensilla and pores presumably homologous to those in Diacyclops ishidai sp. n. Greek letters indicating pores and sensilla homologous to those in Diacyclops hanguk sp. n. Arrows pointing most prominent specific features. Scale bars 100 μm.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.
-
All Biocode files are based on field identifications to the best of the researcher’s ability at the time.